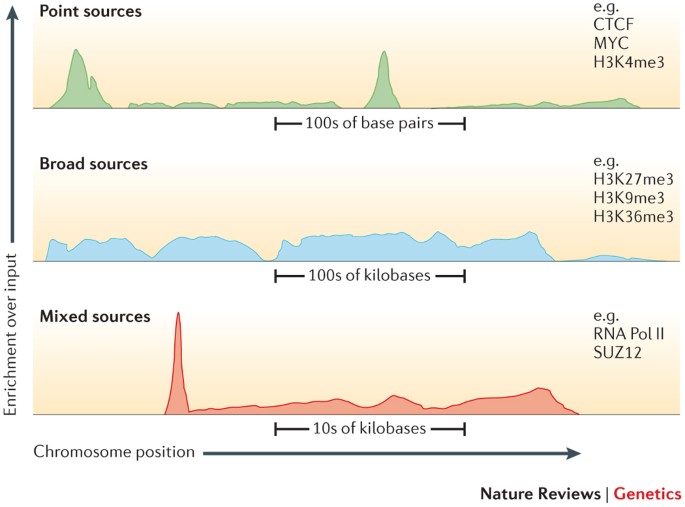
GitHub - GenomicaMicrob/coverage_calculator: A simple script to calculate the coverage of a genome assembly
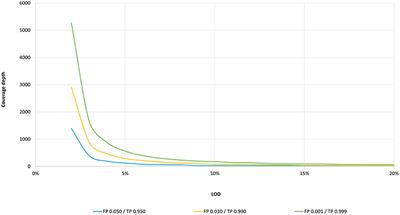
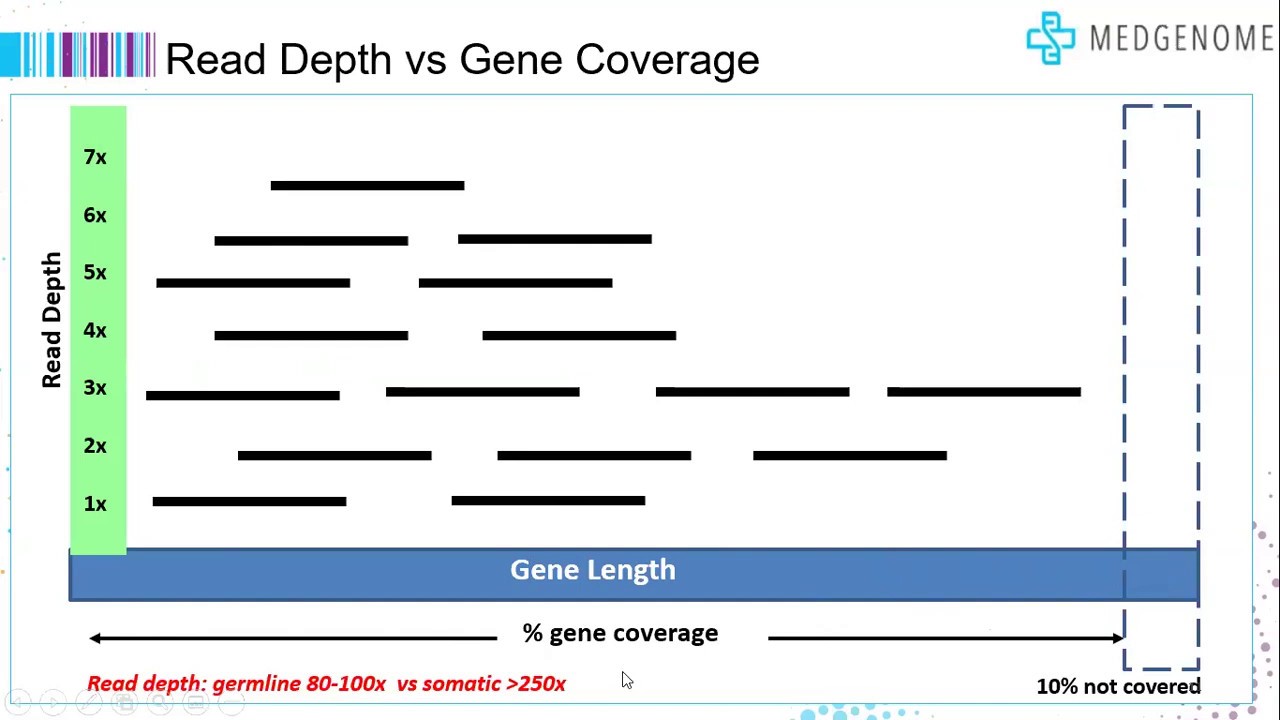
Frontiers | Standardization of Sequencing Coverage Depth in NGS: Recommendation for Detection of Clonal and Subclonal Mutations in Cancer Diagnostics

Covcalc: Shiny App for Calculating Coverage Depth or Read Counts for Sequencing Experiments | R-bloggers

How is the percentage of protein sequence coverage calculated in the search report of MS/MS Ions Search in Mascot? | ResearchGate

A beginner's guide to low‐coverage whole genome sequencing for population genomics - Lou - 2021 - Molecular Ecology - Wiley Online Library
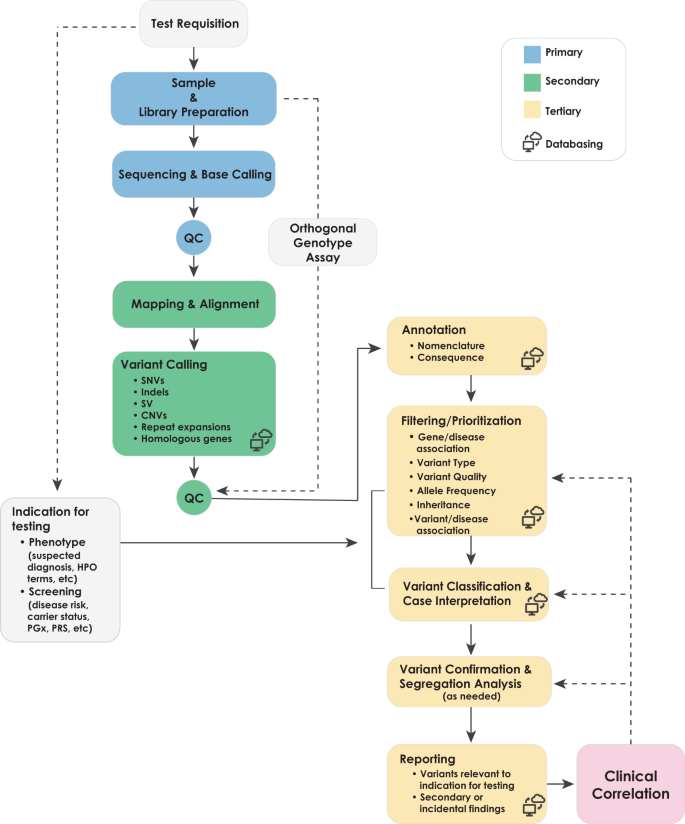
Best practices for the analytical validation of clinical whole-genome sequencing intended for the diagnosis of germline disease | npj Genomic Medicine